Developing a Deep Learning Model for Assisted Pathology Diagnosis
By Ong Kok Haur, Laurent Gole, Huo Xinmi, Li Longjie, Lu Haoda, Yuen Cheng Xiang, Aisha Peng, and Yu Weimiao, Bioinformatics Institute
|
The ready-to-use functions that MATLAB provides reduced the time and effort required for us to develop and implement the platform, enabling us to accelerate our research and focus on generating new insights. |
||
As the second most common form of the disease among males, prostate cancer is typically diagnosed via the careful inspection of tissue samples. This inspection, conducted by expert pathologists using a microscope, is labor-intensive and time-consuming. Additionally, the number of medical professionals capable of doing this work is limited—especially when clinical workloads are high—which can lead to backlogs of samples that must be analyzed and delays in starting treatment.
Emerging in part due to the limitations of manually analyzing samples, research into the use of deep learning to assist with pathology diagnosis for prostate and other forms of cancer has rapidly expanded. Several technical hurdles need to be cleared, however, before deep learning models can be created, trained, and deployed for clinical applications. For instance, about 15% of digital pathology images have quality problems related to focus, saturation, and artifacts, among other issues. Moreover, image quality cannot be quantitatively assessed with the naked eye, and the whole-slide image (WSI) scanners used today produce extremely large data sets, which can complicate image processing with high-resolution images of 85,000 × 40,000 pixels or more. Additionally, like manual pathology diagnosis, the process of annotating images requires a significant amount of time from experienced pathologists, making it difficult to assemble the high-quality database of labeled images needed to train an accurate deep learning model.
Our team at Bioinformatics Institute, an Agency for Science, Technology and Research (A*STAR) Institute in Singapore, has developed a cloud-based automation platform that addresses many of the challenges associated with deep learning–assisted pathology diagnosis, while also reducing the burden on pathologists for image labeling and clinical diagnoses (Figure 1). This platform includes A!MagQC, a fully automated image quality assessment tool developed in MATLAB® with Deep Learning Toolbox™ and Image Processing Toolbox™. The platform also includes a deep learning classification model trained in MATLAB to identify patterns associated with prostate cancer, with pathologists using A!HistoClouds—a Java®-based user interface—to label images and manage inference results. In experiments with pathologists in Singapore and China, the platform reduced image labeling time by 60% when compared to manual annotation and traditional microscopic examination, helping pathologists analyze images 43% faster while maintaining the same accuracy as conventional microscopic examination. The ready-to-use functions that MATLAB provides reduced the time and effort required for us to develop and implement the platform, enabling us to accelerate our research and focus on generating new insights.
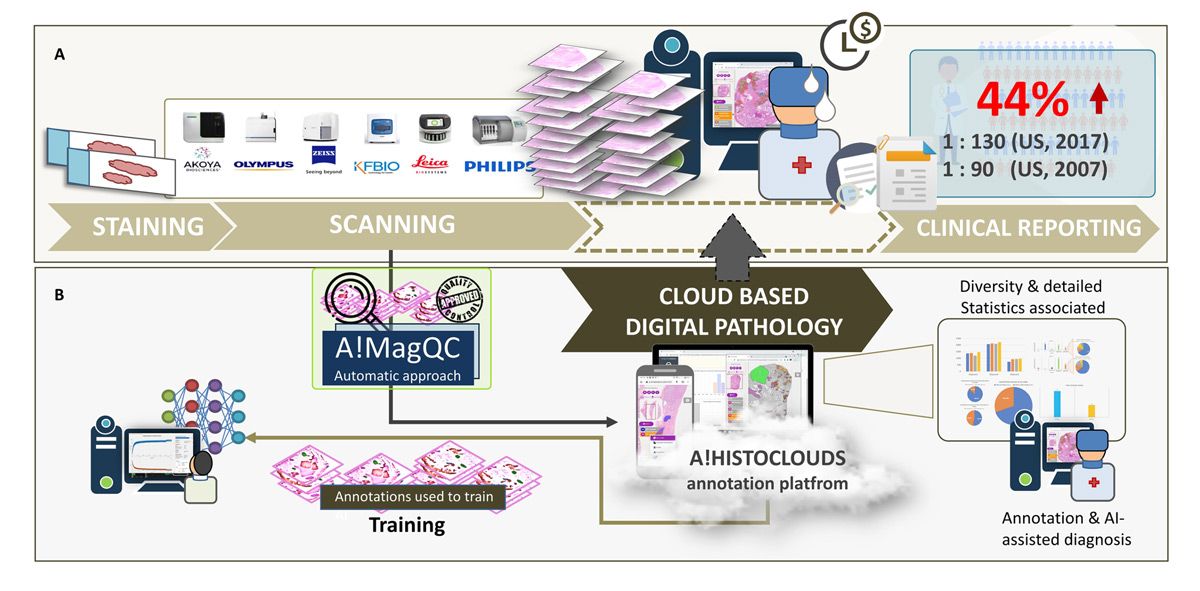
Figure 1. The digital pathology image analysis platform, including A!MagQC and A!HistoClouds.
Image Quality Assessment with A!MagQC
In digital pathology, image quality problems can be broadly divided into two categories: tissue sample preparation problems and scanning problems (Figure 2). Tissue tears, folds, air bubbles, overstaining, and understaining fall into the first category; when these issues are detected a new sample will need to be cut. On the other hand, when image contrast, saturation, and focus problems are detected, the existing sample can simply be rescanned, and recutting is not necessary.
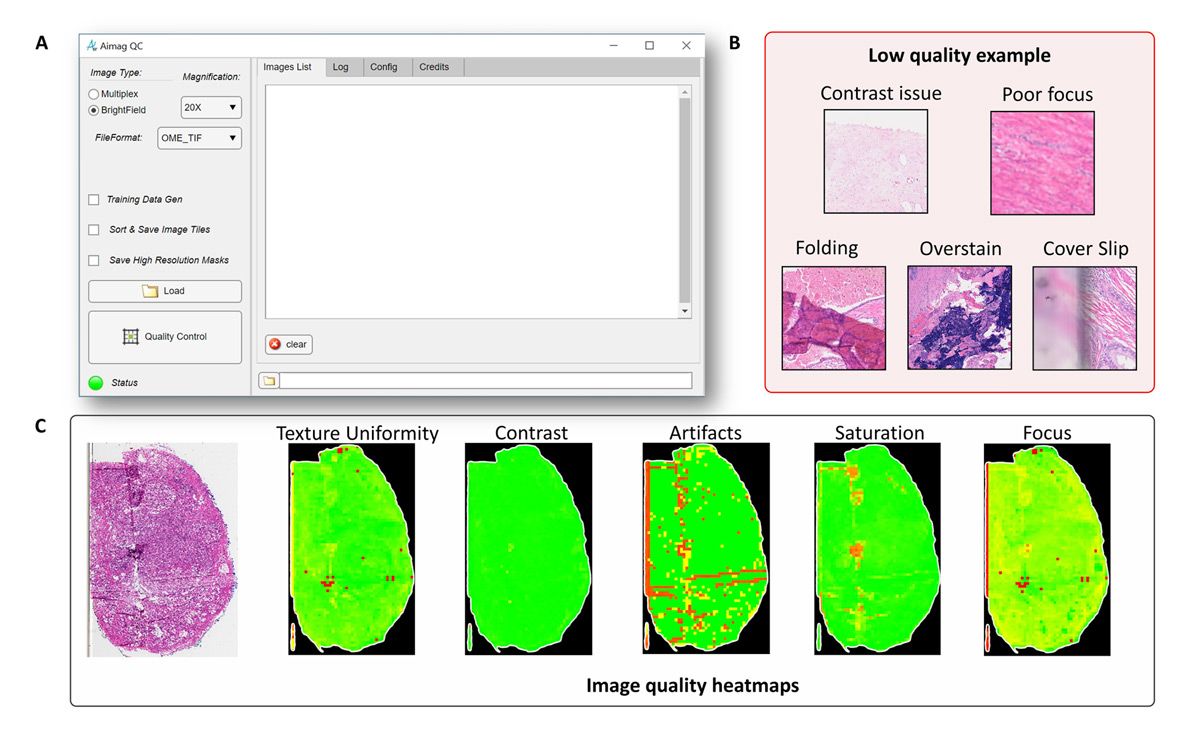
Figure 2. Texture uniformity, contrast, artifacts, saturation, and focus problems detected with A!MagQC.
Whether the analysis is conducted by pathologists or via deep learning models, any of these common problems can have an adverse effect. As such, we developed image processing algorithms in A!MagQC to automatically detect the principal factors affecting image quality. We considered using Python® for A!MagQC development, but chose MATLAB because of the specialized toolboxes and capabilities it offers. Although Python libraries are available for image processing and other subject areas relevant to our work, the dedicated functionality and example applications (including digital pathology applications) that MATLAB provides enabled us to work more efficiently. For example, with images too large to load into memory, we used the blockproc function from Image Processing Toolbox to divide each image into blocks of a specified size, process them one block at a time, and then assemble the results into an output image.
Other capabilities that were particularly useful for us included MATLAB App Designer and MATLAB Compiler™, which we used to build the A!MagQC user interface and to compile our MATLAB code into a standalone A!MagQC executable for distribution, respectively.
Using the finished A!MagQC application, we tested image quality from five different WSI scanners—each from a different vendor—to identify variances in color, brightness, and contrast for the same sample. This exercise helped us ensure that the deep learning model we subsequently trained would produce accurate results for the broad range of scanners in use today.
Model Training and Testing
When analyzing a sample, pathologists apply the Gleason grading system—which is specific to prostate cancer—to assign a score based upon its appearance. Aside from normal or benign tissue, areas of the sample may include stroma (connective tissue) or tissue that is assigned a Gleason score from 1 to 5, with 5 being the most abnormal (Figure 3). Before we could begin training a deep learning model to classify tissue samples, we needed to assemble a data set of image patches that was labeled with these categories. This task was completed with the help of a team of pathologists using A!HistoClouds, who worked with images that had been checked for quality using A!MagQC. Once we had a base set of labeled image patches, we performed data augmentation to expand the training set by reflecting individual images vertically or horizontally and rotating them by a randomly selected number of degrees.
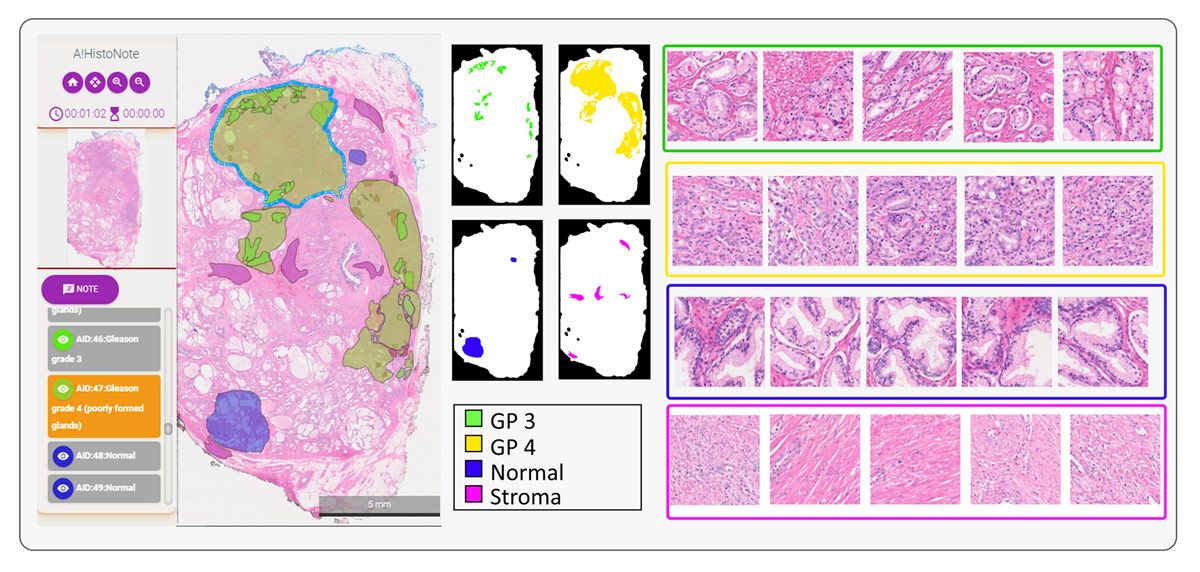
Figure 3. Tissue samples showing stroma, benign tissue, and tissue scored as Gleason 3, Gleason 4, and Gleason 5.
Working in MATLAB with Deep Learning Toolbox, we created our deep learning model structure using ResNet-50, VGG-16, and NasNet-Mobile pretrained networks, replacing their regular classification layers with a weighted classification one (Figure 4). We used Parallel Computing Toolbox™ to make use of the multiple NVIDIA® GPUs installed in our server and sped up training speed by approximately four times. Additionally, scaling from a single GPU card to multiple GPUs is as simple as using the multi-GPU option. We found that ResNet-50 and VGG-16 classifications were more accurate than those of NasNet-Mobile for our data set, and ultimately chose ResNet-50 because of its smaller size.
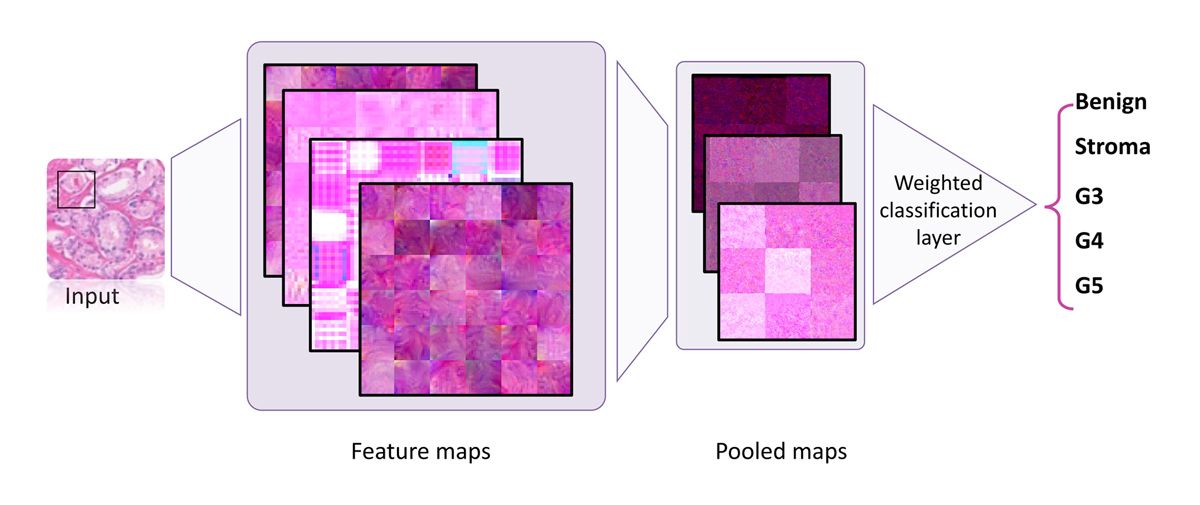
Figure 4. Training structure using a weighted classification layer.
The model is trained and used via an iterative process. Following the first stage of initial training on manually labeled images is a second, semiautomated stage in which pathologists review and modify predictions generated by the trained model (Figure 5). This second stage is repeated until the model is ready to be used by medical professionals to assist with clinical diagnoses.
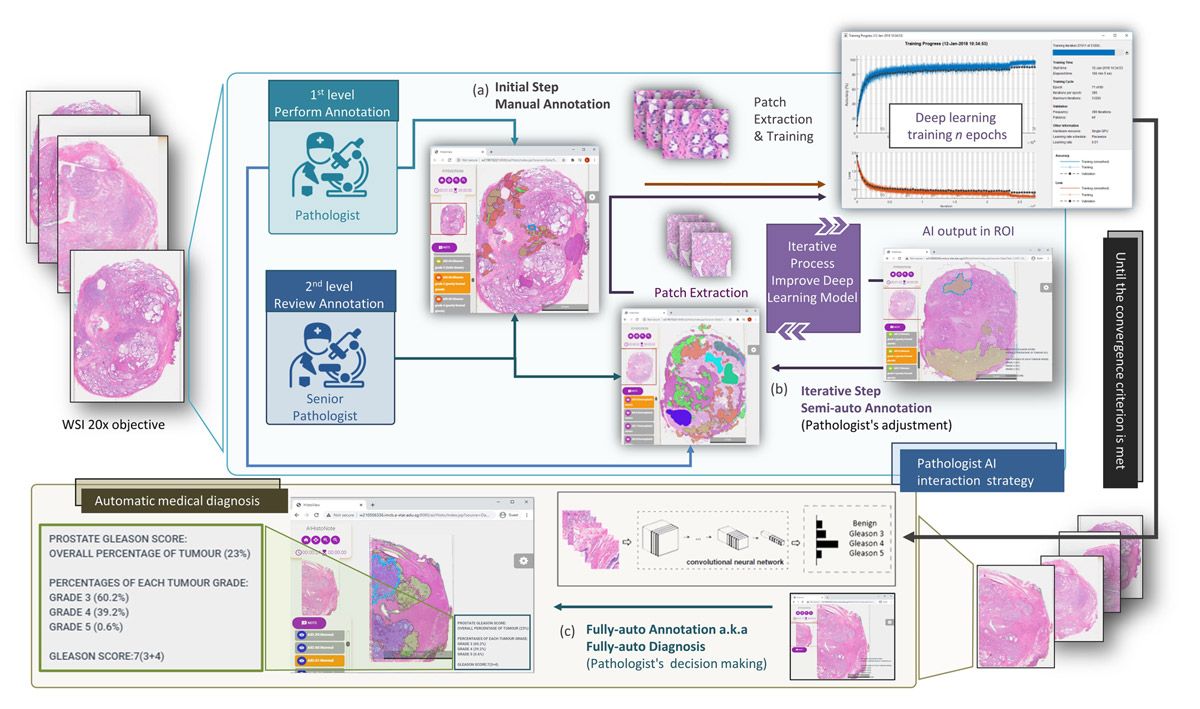
Figure 5. Iterative process for training.
Next Steps
We have deployed our deep learning–assisted pathology diagnosis platform to Amazon® and Huawei Cloud Web Services, providing easy access to a team of pathologists working in different countries. We are continuing to extend the platform’s capabilities, including tighter integration with A!MagQC and a new application, A!Prostate, which is designed for automated Gleason grading on prostate pathological images from diverse sources. We are also working on validating and optimizing our deep learning model for additional clinical scenarios, including different tissue thicknesses, staining mechanisms, and image scanners. Lastly, we are exploring opportunities to extend the same image quality assessment and deep learning workflow beyond prostate cancer to other types of cancer.
We also organized the Automated Gleason Grading Challenge 2022 (AGGC 2022), which was accepted into the 2022 International Conference on Medical Image Computing and Computer Assisted Intervention. AGGC 2022 focuses on addressing challenges in Gleason grading for prostate cancer, digital pathology leverage, and deep learning approaches. The challenge aims to develop automated algorithms with high accuracy for H&E-stained prostate histopathological images obtained from various digital scanners. Notably, this is the inaugural challenge in the field of digital pathology that investigates variations in color induced by image scanning processes.
Although the challenge has concluded, the complete data set is now available for continued research.
Published 2024
Products Used
- MATLAB
- Deep Learning Toolbox
- Image Processing Toolbox
- MATLAB Compiler
- Parallel Computing Toolbox
Download a FREE Trial Request Consultation